Main menu
You are here
Model entry
| Search | Simulistics | Model catalogue | Listed by keyword | Listed by ID | Listed by title | Listed by date added |
Conway's Game of Life
Model : Life
Simile version : 4.0
Date added : 2004-10-19
Keywords :
One sided enumeration ;
Cellular automata ;
Description
Conway's Game of Life
This is just one example of a cellular automaton, which is any system in which rules are applied to cells and their neighbors in a regular grid. See the Wonders of Math site for a detailed description of the game and its history.It was invented by mathematician John Conway in 1970 as an illustration of how complexity can emerge from simple starting conditions. Writing an implementation of the Game of Life is often set as an exercise for students, and it is included here to illustrate Simile's capabilities as a graphics-based programming language, and in particular the ability to set the membership of an association submodel in which one base model contains the information needed to choose the instances of the other base model with which they are associated. This is a new feature in Simile 4.0.
How to run it
To set an initial pattern, go to the 'start pattern editor' in the execution window, zoom in to the grid a bit and select the 'Enter edit mode' icon in its menubar. Now click on the colour scale near the top end, and click on any squares you want to be live at the start. When the model is next reset, this pattern will appear in the main grid.
How it works
The model is simpler than it looks. The stuff to the left hand side is all dedicated to allowing the initial state to be set by means of a small grid which is displayed and edited with the polygon I/O tool. This grid is mapped onto the middle of the main grid, allowing simple patterns to be entered which then evolve into larger more complex ones.This version of the game does not wrap around at the borders; squares beyond the edge are treated as never being alive. This means that not all squares in the grid have the same number of neighbours. Those in the middle have eight, those on the edges have five and those in the corners have three. This is taken care of by making a variable-membership submodel for neighbours within the cell submodel. The membership does not vary with time, it just varies between cell instances.
In the neighbours submodel, the existence condition checks whether each neighbour falls within the grid. If it does, it calculates the neighbour's index by reversing the process of calculating row and column from the index. So the output of this submodel is a list of each cell's neighbours' indices.
Now for the clever bit. The 'test' relation arrow is flagged to allow base instance lookup, which ensures that its index will be index(1) in the relation model. The expression for the relation's existence condition is
any(index(1) is {neighbours_ids_env}). This
produces the same behaviour as any(index(1) ==
{neighbours_ids_env}) but is more efficient. If the equality
test is used, the program Simile builds will create a loop to get the
list of neighbours from each instance of cell, and another loop nested
within that to look at the indices of all the cells and create a
relation instance between it and the first cell if its index appears
in the first cell's list of neighbours' indices. With 40,000 cells
this means doing 800 million checks. However, using 'is' instead of
'==' specifies one sided relation enumeration. In this case the outer
loop is the same, but for each instance the relation instances are
created directly with all the cells whose indices are in the neighbour
list, rather than by searching through them again.
Having created the relation for neighbourhood, it is simple for each cell to sum the number of live neighbours, allowing it to work out whether it is born, survives or dies. Running the model is fast, because the relations only have to be established when it is initialized. Unfortunately the grid display tool takes about a second to update with this many grid squares, so it works best with the display interval set to 10 or so model time steps.
Files
Model fileClick on the icon to download the model file. (You will need Simile to examine and run the model. A free evaluation version is available from the products page.) Some browsers may attempt to display the model file, rather than open it in Simile; in this case, use the browser back button to return to this page, and use the context menu (invoked by right-clicking on the link) to save the target file to disk. |
|
|
|
|
|
|
Saved display tool state for Life model |
Diagram
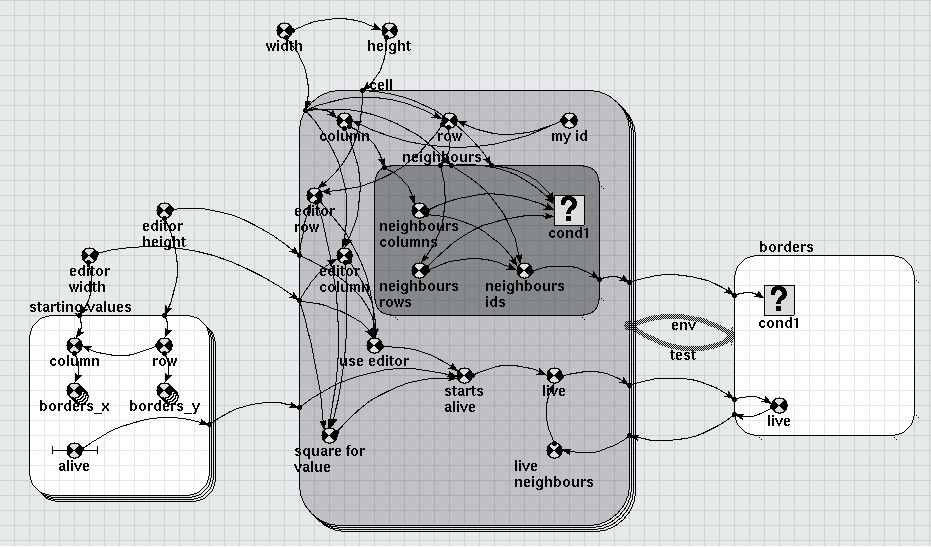
Results
|Here is the display 100 generations into the 'puffer train' example from the Wonder of Math site |
|Web links
Wonders of MathReferences
Bibliography
Berlekamp, Conway, and Guy: Winning Ways (for your Mathematical
Plays), Volume 2, (c)1982. ISBN 0-12-091152-3.
Gardner, Martin: Wheels, Life, and Other Mathematical Amusements,
(c)1983. ISBN 0-7167-1589-9.
Poundstone, William: The Recursive Universe, (c)1985. ISBN
0-688-03975-8.
Copyright 2003-2026 Simulistics Ltd
